Rise of the molecular machines
Chemists are putting molecules to work in ways only limited by our imagination
Josh Howgego
Our bodies are full of molecular machines and their synthetic counterparts may soon be all around us too
© SCIENCE PHOTO LIBRARY
|
Nuts and bolts
Mastering and mimicking how molecular machines work has been a fascinating adventure for chemists, and one in which their creativity has been unleashed. The parts that make up the tiny world of molecular machines are described using familiar terms: an alkyl chain might be called a piston, for example, or a benzene ring a wheel. But before these tiny machines could be built, an in depth knowledge of how to hold the component pieces together was needed.
Instead of nuts and bolts, molecular machines are held together by intermolecular forces; the subtle electrostatic attractions and repulsions between molecules. Chemistry is inherently dynamic, so these forces can hold molecules together strongly, but also enforce softer attractions, which pull molecules together, but still allow some degree of movement (see table.1 And it is these softer interactions that are key to many molecular machines.
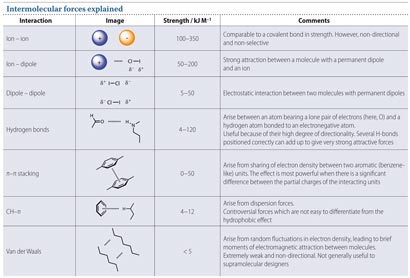
Intermolecular forces explained. Download the pdf at the end of the article to see a high resolution version of this image.
|
Indeed one of the main steps towards the first molecular machine was working out how to mechanically interlock two molecules, whilst still allowing them freedom to move; a bit like having a ring slide along a piece of string. This problem was pondered for a long time, until the 1990s when several developments converged and made it possible.
A molecular shuttle
Fraser Stoddart, a chemist at Northwestern University, US, produced one of the early prototype molecular machines in 1994 (fig?1).2 The machine has a track with two stations and a shuttle that can slide along between them. The shuttle can't escape from the end of the track because there are bulky stopper groups at both ends which it can't fit over.
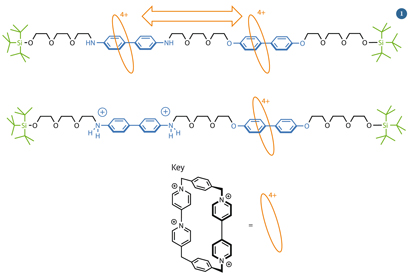
Fig 1 - The key parts of Stoddart's molecular shuttle are the two stations (blue), the shuttle (orange) and the bulky stoppers (green)
|
The intermolecular forces that exist between the shuttle and the different points on the track determine where it sits. Because the shuttle is made from non-polar, aromatic building blocks, the  -
- stacking intermolecular forces between it and the aromatic ring-based stations are quite strong. The forces between the shuttle and the track made of polyether chains are much weaker, as there are no aromatic moieties to stack against.
stacking intermolecular forces between it and the aromatic ring-based stations are quite strong. The forces between the shuttle and the track made of polyether chains are much weaker, as there are no aromatic moieties to stack against.
Under normal conditions the shuttle doesn't have much preference between the two stations, so it is free to move rapidly over the polyether chain and between the two ports. However, the machine is cleverly designed to change its properties under different conditions. The nitrogen atoms in the first station are basic. That is, they are easily able to accept a hydrogen ion (proton) on exposure to acid. In the process the first station gains a positive charge. This repels the shuttle (which also has a positive charge), and so the shuttle is pushed to the second station.
What the chemists had made was essentially a switch. With the addition of a little acid, their machine switched from freely moving between both states to being confined at a single station.
Molecular muscles
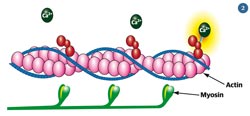
Fig 2 - The actin and myosin filaments in muscles use protein-based machines to pull past each other
|
The ability to emulate the molecular machines which power logistics operations in living systems is also desirable. Before mitosis, chromosomes need to be copied, and then each set must be transported to opposite ends of their cell. The biological solution is a network of microscopic fibres called microtubules strung across a cell's diameter, and kinesin (a motor protein) that can move along these rails. The kinesin hitches up its chromosone cargo, and hauls it across the cell.
But so far molecular machines have been employed most successfully as information storage media (the 'freely moving' and 'stuck' modes of the shuttle can be interpreted as a digital readout: off or on; 0 or 1). In contrast, the challenge of making artificial muscles or kinesin mimics is hard because one molecular machine on its own is too weak to be useful. Transferring controlled motion from a single interlocked molecule to a complex system - where lots of shuttles work together - has proved difficult.
Some prototype molecular machines have been developed, however, which - although drastically simplified - work similarly to the actin and myosin in muscles. But these don't use intermolecular forces in the same way as Stoddart's shuttle. One model machine, developed by chemist David Leigh, at the University of Edinburgh, UK, instead works by sequentially making and breaking different types of covalent chemical bonds.3
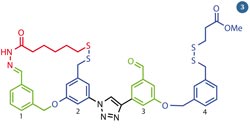
Fig 3 - Leigh's molecular walker can take steps along a pathway of alternating functional groups
|
Like Stoddart's machine, the impetus for the walker's motion comes from varying the nature of the solution surrounding the molecules from acidic to basic. To start the walker off, Leigh uses acidic conditions. This fixes the disulfide (S-S) linkage in place, but means the imide (N-N) bond is under equilibrium; it is constantly breaking and re-forming. When the imine bond is broken, the walker can pivot on its disulfide foot, and the imine foot is free to either re-form the bond with foothold 1 or move forwards to form a new bond with the aldehyde moiety at foothold 3 . (Importantly, the walker unit must be attached by at least one 'foot' at all times. It couldn't run or jump, as with both linkages broken the walker would drift off into solution).
By increasing the pH of the solution, the walker is switched from having its disulfide foot fixed to the track to having the imine foot held down. The basic conditions mean that the disulphide bond is under equilibrium this time, and now it can move around.
Nature generally uses non-covalent interactions to stick the feet of its protein machines to their tracks. So although Leigh's model walker is not strictly biomimetic, it is an ingeniously simple way of achieving
much the same outcome as cellular transporting machines.
much the same outcome as cellular transporting machines.
In the future, walkers operating on extended tracks could be made. And in theory it should be possible to attach cargo to a walker. This would make an excellent analogue of the chromosone-transporting kinesin protein.
Making nanocars
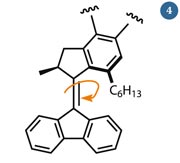
Fig 4 - Feringa's molecular rotor is powered by electrons, which excite the double bond and induce an E/Z-isomerisation
|
Perhaps that is why Ben Feringa, a chemist working at the University of Groningen in the Netherlands, also steered clear of them. His interest lay in making machines that produce rotary, rather than linear, motion. His rotary motor (fig 4) is composed of a single part, allowing him to avoid sticking axles and levers together using intermolecular forces.
Feringa recently combined four of his rotary motors with a chassis to develop what he called a four-wheeled molecule (but everyone else likes to call a nanocar).4 In order for the car to drive along a surface, the paddle-like rotors must turn. The surface the car travels along is a metal, with electricity running through it. The electrons excite the rotor's double bond and induce an E/Z -isomerisation. The steric strain then leads to a helix inversion, and the molecule flips over, producing rotational motion and powering the car forwards (fig?5).
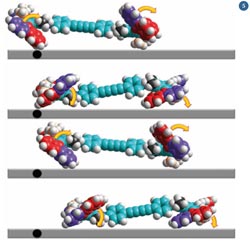
Fig 5 - If the four rotors work in unison, the nanocar begins to drive along the surface
|
Today, molecular machines that can perform an increasingly varied range of tasks are still being constructed. Chemists have succeeded in replicating the sorts of machines nature uses in our bodies with artificial systems in single interlocked systems. These machines are truly biomimetic: they operate using the same principles that nature has used since its inception. Impressive, given that it took many millennia for them to evolve in the first place.
The Nobel prize winning scientist Richard Feynman talked about molecular machines in 1959, when people had just begun to imagine what they might be like. 'What would be the utility of such machines?' he said. 'Who knows? I cannot see exactly what would happen, but I can hardly doubt that when we have some control over the arrangement of things on a molecular scale we will get an enormously greater range of possible properties that substances can have, and of the different things we can do.'
What exactly the legacy of molecular machines will be still remains to be seen, but with molecular shuttles and nanocars already behind us, we can safely agree with Feynman that the possibilities are exciting.
Josh Howgego is a science writer and chemistry PhD student based in Bristol, UK
References
1 Bond strengths are taken from: J W Steed and J L Atwood, Supramolecular chemistry, Wiley, 2000; M Nishio et al, CrystEngComm., 2009, 11, 1757 (DOI:10.1039/b902318f)
2 R A Bissell et al, Nature, 1994, 369, 133 (DOI:?10.1038/369133a0)
3 M von Delius, E M Geertsema and D A Leigh, Nat. Chem., 2010, 2, 96 (DOI:?10.1038/nchem.481)
4 T Kudernac et al, Nature, 2011, 479, 208 (DOI:?10.1038/nature10587)
2 R A Bissell et al, Nature, 1994, 369, 133 (DOI:?10.1038/369133a0)
3 M von Delius, E M Geertsema and D A Leigh, Nat. Chem., 2010, 2, 96 (DOI:?10.1038/nchem.481)
4 T Kudernac et al, Nature, 2011, 479, 208 (DOI:?10.1038/nature10587)
No comments:
Post a Comment